This build of rgl does not include OpenGL functions. Use
rglwidget() to display results, e.g. via options(rgl.printRglwidget = TRUE).null
1 

Author: Dr. Hamed Abdollahi
PI: Dr. Homayoun Valafar
Vectors are single-dimensional, homogeneous data structures.
So, Vectors can only contain elements of one type!
To create a vector, concatenating values using the c() function.
This build of rgl does not include OpenGL functions. Use
rglwidget() to display results, e.g. via options(rgl.printRglwidget = TRUE).null
1 Vectors must have their values all of the same mode.
In vectors, cells are accessible through indexing operations.
Some important functions are: c() , vctor(),length(), class(), typeof(), attributes(), is.double(), is.numeric(), is.integer(), is.logical(), is.character(), is.vector() .
A vector can be empty and still have a mode.
For example, an empty character string vector is represented as character(0), while an empty numeric vector is represented as numeric(0).
Numeric sequences:
Regular sequences can be created using the function seq() and its shortcuts seq.int(), seq_along(), and seq_len().
numeric or integer values between two well defined points (from and to) with an equidistant spacing.
LETTERS and lettersType of vector in R used to represent categorical data where the levels of the factor represent distinct groups or categories.
The tapply() function in R is used to apply a function (such as mean(), sum(), median(), etc.) to subsets of data defined by factors or groupings. It allows you to compute a statistic for each group defined by levels of a factor, treating them as separate entities.
 Let’s start Exercise_4!
Let’s start Exercise_4!
 We delved into creating projects and utilizing the icons on the toolbar to streamline our workflow.
We delved into creating projects and utilizing the icons on the toolbar to streamline our workflow.
 We learned how to check for package issues using the rcmdcheck package, ensuring our packages function correctly.
We learned how to check for package issues using the rcmdcheck package, ensuring our packages function correctly.
 We discussed various AI-APIs and packages related to R, exploring their applications and integrations.
We discussed various AI-APIs and packages related to R, exploring their applications and integrations.
 We practiced retrieving data from the PDB API, gaining hands-on experience in fetching and handling external data sources.
We practiced retrieving data from the PDB API, gaining hands-on experience in fetching and handling external data sources.
 We had a brief introduction to OOP concepts and how they apply within the R programming language.
We had a brief introduction to OOP concepts and how they apply within the R programming language.
 We discussed the structure and architecture of R, focusing on classes and objects, understanding how R is built and how it operates.
We discussed the structure and architecture of R, focusing on classes and objects, understanding how R is built and how it operates.
 We became familiar with attributes and methods in R, as well as the vector data type.
We became familiar with attributes and methods in R, as well as the vector data type.
Types of Factors:
ordered() function creates such ordered factorsImportance of factors:
Replicating elements
Replicate one specific value n times.
Given an existing vector: Replicate each element n times.
Given an existing vector: Replicate the entire vector n times.
Given an existing vector: Replicate the elements different amount of times.
he function rep.int() in R is used to replicate elements of a vector or a list.
To find the sample size (number of observations) in R using the length() :
If we combine elements of different types, R has to convert all elements into the same type/class as vectors can only contain elements of one type.
This is called ‘coercion’. So, In R, coercion occurs when elements of different types are combined in a vector:
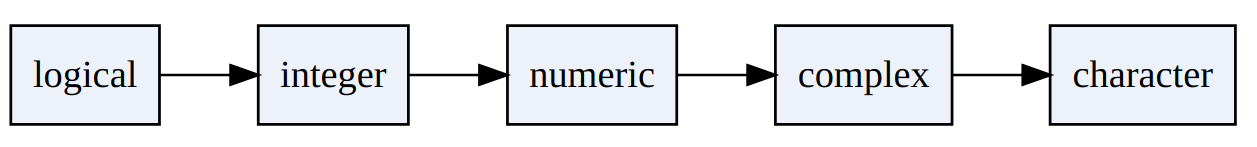
Every numeric value equal to 0/0L converted to logical results in FALSE.
Every numeric value not equal to 0/0L converted to logical results in TRUE.
Every TRUE converted to numeric will be 1 (or 1L).
Every FALSE converted to numeric will be 0 (or 0L).
Explicit coercion: we force something to be of a different type:
as.integer() , as.numeric() ,as.character() ,as.logical() ,as.matrix().
If R is not able to convert elements, it will return NA
Multiply a sequence with a scalar (a single number)
How R handles vector arithmetic:
Negative Indices: Negative indices can be used to exclude specific elements.
Logical Indices: Logical vectors can also be used for indexing, selecting elements based on conditions.(Select elements based on logical conditions TRUE or FALSE)
A matrix is a two-dimensional data structure where elements are organized into rows and columns.
It is homogeneous, meaning all elements within a matrix must be of the same data type (e.g., numeric, character, logical).
null
3 If different data types are attempted to be combined into a matrix using matrix(), R will coerce them into a common type that can accommodate all elements.
Matrices are created using the matrix() function, which takes vectors as input and arranges them into a specified number of rows and columns.
The %o% operator in R is used to compute the outer product of two vectors.
The outer product ab will be a matrix where each element ab[i,j] is a[i]×b[j].
Matrix Transpose (t()):
The t() function computes the transpose of a matrix. It swaps rows with columns.
An array is a multiply subscripted collection of data entries, typically of the same data type, such as numeric values.
Arrays are generalizations of matrices and can have multiple dimensions. A way to store and manipulate multi-dimensional data beyond the two dimensions provided by matrices.
Arrays can have multiple dimensions.
Dimensions in R arrays are indexed from one up to the values given in the dimension vector. This means the first dimension of an array is indexed by 1, the second by 2, and so on.
null
5 Using Vectors as Arrays:
In R, a vector can be treated as an array if it has a dim attribute set. This allows you to reshape a vector into a multi-dimensional array using the dim() function or directly assigning the dim attribute.
When working with arrays and matrices:
c(): The c() function in R concatenates its arguments to create a single vector. However, when used with arrays or matrices, c() disregards any dimension attributes (dim and dimnames). This means it treats the input as a flat sequence of elements and does not respect the structure of arrays or matrices.Difference from cbind() and rbind(): Unlike cbind() (column bind) and rbind() (row bind), which respect the dim attributes of matrices and arrays, c() does not preserve these attributes. This behavior can be useful when you specifically want to flatten an array or matrix into a vector format.
Coercion to Vector: To convert an array or matrix back to a simple vector while preserving its structure, the recommended approach is to use as.vector(). For example:
This function maintains the structure of the object, ensuring that it remains a vector but retains any dimension attributes.
c
Lists can contain objects of different types and structures.
Lists have elements, each of which can contain any type of R object.
Warning in brewer.pal(n = 16, name = "Dark2"): n too large, allowed maximum for palette Dark2 is 8
Returning the palette you asked for with that many colorsnull
7 There is no particular need for the components to be of the same mode or type, or example, a list could consist of a numeric vector, a logical value, a matrix, a complex vector, a character array, a function, and so on.
If Lst is the name of a list with four components, these may be individually referred to as Lst[[1]], Lst[[2]], Lst[[3]] and Lst[[4]].
If, further, Lst[[4]] is a vector subscripted array then Lst[[4]][1] is its first entry.
list().The basis for most data analyses in R are data frames.
Data frames are indeed structured as lists with class “data.frame”.
Data frames are widely used in R for storing and manipulating structured data,
Data frames have specific rules regarding their composition and structure:
Variables in Data Frames: Matrices, lists, and data frames contribute variables (columns) to the new data frame based on their columns, elements, or variables, respectively.
Each column represents a variable (e.g., age) and each row represents an observation (e.g., an individual).
Consistent Length and Size: Variables (columns) within a data frame must have consistent lengths for vectors and consistent row counts for matrices. This ensures uniformity across columns in terms of data structure.
Matrix-Like Operations: While data frames are list-like structures, they can be treated like matrices in many operations. They can be displayed in matrix form, and their rows and columns can be accessed using matrix indexing conventions.
null
9 Language objects can be converted to and from lists by the as.list and as.call functions.
Symbols can be created through the functions as.name and quote.
The main difference between language object and expression object is that an expression object can contain several such expressions.
tibble or tbl_df is a modernized version of data frame, provided by the tibble package, designed to address some of the shortcomings of traditional data frames.
They maintain compatibility with data frames but offer enhanced features and better printing defaults. Here’s how you can use as_tibble() to coerce objects into tibbles:
Key Characteristics of Tibbles:
Lazy Evaluation: Tibbles are lazy, meaning they do not automatically change variable names or types. This characteristic encourages explicit and intentional programming practices, minimizing unexpected changes and errors.
Surly Behavior: Tibbles complain more assertively than data frames. For example, if you attempt to access a variable that doesn’t exist, a tibble will raise an error, prompting you to address issues earlier in your code development process.
Enhanced Print Method: Tibbles feature an enhanced printing method that improves readability and usability, especially with large datasets containing complex objects. This makes it easier to interact with and understand data directly from the console or within scripts.
Simpler Coercion: The as_tibble() function simplifies the process of coercing objects into tibbles compared to as.data.frame() methods. This simplification enhances code clarity and reduces the cognitive load when working with different data structures.
Tibbles do not support row names. They are removed when converting to a tibble or when subsetting:
Benefits of Using Tibbles:
Cleaner Code: By enforcing stricter rules and providing clearer error messages, tibbles help maintain cleaner and more expressive code.
Improved Debugging: Early error reporting and clear feedback on problematic operations facilitate quicker debugging and troubleshooting.
Compatibility with Modern Data Analysis Tools: Tibbles are designed to integrate seamlessly with modern data analysis tools and packages in R, supporting efficient and effective data manipulation and visualization tasks.
Recycling
When constructing a tibble, only values of length 1 are recycled. The first column with length different to one determines the number of rows in the tibble, conflicts lead to an error:
| Feature | Matrix | DataFrame | DataTable | Tibble |
|---|---|---|---|---|
| Column Types | Homogeneous | Heterogeneous | Heterogeneous | Heterogeneous |
| Memory Efficiency | High | Moderate | High | Moderate |
| Computation Speed | Fast | Moderate | Very Fast | Moderate |
| Ease of Use for Data Manipulation | Low | High | High | High |
| Integration with tidyverse | No | Partial | Partial | Yes |
| Learning Curve | Low | Low | High | Moderate |
Missing values in R still have a class.(missing numeric, integer, logical, or character missing values.)
NaN: Mathematically not defined (and always of class numeric).
NA: Missing value, NA’s still have classes!
NaN is also NA but not vice versa.
Control structures in R allow you to manage the flow of execution for a series of expressions.
Control flow in R involves managing how code execution proceeds based on conditions and iterations.
They enable you to add logic to your R code, making it more dynamic and responsive to different inputs or data features.
By using control structures, you can ensure that different R expressions are executed based on certain conditions, rather than always running the same code each time.
They are typically used in functions or longer expressions rather than in interactive sessions.
Here are some commonly used control structures:
for loop: Used to execute a loop a specified number of times.
break : Used to terminate the execution of a loop prematurely.
next : Used to skip the current iteration of a loop and proceed to the next iteration.
while : Used to execute a loop as long as a condition remains true.
if and else : Used to test a condition and execute code based on the result.
repeat :Used to execute an infinite loop, which must be explicitly terminated with a break statement.
There are two type of control flow:
Choices (Conditional statements)
Loops (Iteration)
Invalid Inputs
The condition should evaluate to a single TRUE or FALSE.
Handling Missing Values:
ifelse() propagates missing values in its output. If the test condition results in NA, the corresponding output will also be NA
 For example in this code, the
For example in this code, the NA in x results in "Missing" in result .
Output Type Consistency:
ifelse() expects the yes and no arguments to be of the same type or able to be coerced to the same type.
Mixing different types might lead to unexpected results or coercion that may not align with your expectations.
 Here, because
Here, because "High" is a character and 1:4 is numeric, ifelse() coerces the numeric values to characters where the condition is TRUE.
Alternative Considerations:
If you have multiple conditions and need to handle various types or more complex scenarios, consider using case_when() from dplyr:

case_when() offers more flexibility in handling multiple conditions and provides clearer syntax compared to nested ifelse() statement.
switch() statementThe switch() statement in R is a powerful control structure for handling multiple conditions more compactly than a series of if-else statements.
It allows you to select one of several possible actions based on the value of an expression.
When using switch(), it’s important to ensure that unmatched inputs are handled properly to avoid unexpected NULL returns.
```{webr-r}
# Define a function using switch() to handle multiple conditions
action <- function(option) {
switch(option,
"add" = {
result <- 1 + 1
paste("Adding: 1 + 1 =", result)
},
"subtract" = {
result <- 2 - 1
paste("Subtracting: 2 - 1 =", result)
},
"multiply" = {
result <- 2 * 2
paste("Multiplying: 2 * 2 =", result)
},
"divide" = {
result <- 4 / 2
paste("Dividing: 4 / 2 =", result)
},
stop("Invalid option. Please choose one of 'add', 'subtract', 'multiply', 'divide'.")
)
}
# Example usage
print(action("add")) # Outputs: "Adding: 1 + 1 = 2"
print(action("subtract")) # Outputs: "Subtracting: 2 - 1 = 1"
print(action("multiply")) # Outputs: "Multiplying: 2 * 2 = 4"
print(action("divide")) # Outputs: "Dividing: 4 / 2 = 2"
print(action("unknown")) # Throws an error: "Invalid option. Please choose one of 'add', 'subtract', 'multiply', 'divide'."
``` Explanation:
Explanation:
The switch() statement takes a single expression (option in this case) and matches it against several possible cases ("add", "subtract", "multiply", "divide").
For each case, the corresponding block of code is executed.
If none of the cases match, the last component of the switch() statement (stop()) throws an error, ensuring that unmatched inputs do not silently return NULL.
if multiple inputs should result in the same output, you can leave the right-hand side of the = empty for those cases, allowing the input to “fall through” to the next specified value.
R excels at handling repetitive tasks through loops.
Loops allow you to repeat a set of operations multiple times or until a specified condition is met.
There are three main types of loops in R:
The for loop :
For loops are indeed versatile and commonly used in R for iterating over sequences or performing repetitive tasks.
While other loop constructs like while loops and repeat loops have their specific use cases, for loops often suffice in practice.
The while loop
The repeat loop.
In R, for loops are a fundamental construct used to iterate over elements of objects like lists or vectors. For example:
for each character i in the sequence ‘A’ to ‘J’ do // execute code within curly braces end loop
While other looping constructs exist, for loops are typically adequate for most data analysis tasks due to their simplicity and effectiveness.
When iterating over a vector of indices in R, it’s conventional to use short variable names such as i, j, or k.
 The
The for loop assigns the item to the current environment, which can overwrite any existing variable with the same name. This means that if you have a variable named i in your current environment and you use i as the loop variable, the original i will be overwritten.
 Explanation:
Explanation:
The initial value of i is set to 10.
for loop, i is used as the loop variable and takes on the values from the indices vector (1 to 5).After the loop, the value of i in the current environment is the last value assigned in the loop, which is 5, thus overwriting the original value of 10.
To avoid overwriting variables unintentionally, you can use a different variable name or explicitly manage the scope of your variables:
you can terminate a for loop early using either next or break statements:
next exits the current iteration and proceeds to the next iteration of the loop.
 Explanation:
Explanation:
The next statement is used to skip even numbers. When i is even, the next statement is executed, and the loop proceeds to the next iteration without executing the print statement.
As a result, only odd numbers are printed: 1, 3, 5, 7, 9.
break exits the entire for loop immediately.
 Explanation:
Explanation:
The break statement is used to exit the loop when i is greater than 5.
As a result, the loop stops after printing 1, 2, 3, 4, and 5.
When using for loops in R, there are three common pitfalls to be aware of:
Preallocating Output Containers:
If you are generating data inside a loop, it’s crucial to preallocate the output container. Otherwise, the loop will be very slow.
You can use the vector() function to preallocate the output container.
Avoiding 1:length(x):
Using 1:length(x) can fail in unhelpful ways if x has length 0, resulting in errors
This occurs because : works with both increasing and decreasing sequences. Instead, use seq_along(x), which always returns a value the same length as x.
Handling S3 Vectors:
When iterating over S3 vectors, loops typically strip the attributes. To avoid this issue, call [[ yourself to ensure attributes are preserved.
Using seq_along() for safe iteration.
There are two related tools with more flexible specifications:
while(condition) action:
This performs an action while the condition is TRUE.
repeat(action):
This repeats the action forever (i.e., until it encounters break)
R does not have an equivalent to the do {action} while (condition) syntax found in other languages.
You can rewrite any for loop to use while instead, and you can rewrite any while loop to use repeat, but the converses are not true.
while is more flexible than for, and repeat is more flexible than whileYou shouldn’t need to use for loops for data analysis tasks, as map() and apply() functions already provide less flexible solutions to most problems.
for loopsWe can nest for loops inside one another.
This allows you to perform more complex iterations and computations where multiple levels of looping are required.
Nested for loops involve placing one or more for loops inside the body of another for loop.
Each inner loop executes its entire cycle for every iteration of the outer loop.
This nested structure is useful for iterating over multidimensional data structures like matrices or performing repetitive tasks that involve multiple levels of iteration.
Let’s proceed with Exercise 3.
To comprehend functions in R thoroughly, it’s essential to grasp two key concepts:
Function Components: Functions consist of three primary components:
Formals (formals()):
Body (body()):
Environment (environment()):
Primitive Base Functions: Despite the general rule that functions are defined in R, there are exceptions. A subset of functions known as “primitive” base functions is implemented directly in C for efficiency reasons.
Such as sum() as type builtin and [ as type special,which are exceptions to the standard function structure.
They directly call C code for execution and offering optimized performance for basic operations like summing elements or extracting subsets of data.
Hence, when you check their attributes using formals(), body(), or environment() functions, they typically return NULL because these attributes are not applicable to primitive functions
Primitive functions in R are typically found in the base package of R.
Functions as Objects: Similar to vectors and other data types, functions are objects in R.
R does not require special syntax for defining and naming functions.
You create a function object using the function keyword and then bind it to a name using the assignment operator <-.
This flexibility allows for dynamic and powerful programming capabilities within R.
srcref is useful for printing or displaying the original source code that was used to create the function.
Anonymous functions, also known as lambda functions, e useful in situations where you need a function for a short period or when it’s not necessary to assign a name.
lapply along with anonymous functions defined using function(x).Putting functions in a list can be very useful, especially when you need to store multiple functions together for organizational purposes or to pass them as arguments to other functions.
In the above example:
We define three functions (square, cube, and sqrt).
We create a list function_list containing these functions, where each function is assigned a name within the list (square, cube, sqrt).
We then call each function from the list using $ notation (function_list$square, function_list$cube, function_list$sqrt).
If we have the arguments for a function already stored in a data structure, such as a list or a vector, we can still call the function using the do.call() function in R.
Understanding these aspects is fundamental for mastering function usage and manipulation in R.
Base R provides two ways to compose multiple function calls: nesting and piping.
Nesting
It is straightforward but can become unwieldy with complex chains of functions.
In this example:
mean(data) computes the mean of the data.
data - mean(data) computes the deviations from the mean.
(data - mean(data))^2 squares the deviations.
mean((data - mean(data))^2) computes the mean of the squared deviations (variance).
sqrt(mean((data - mean(data))^2)) takes the square root of the variance to get the standard deviation.
Using a Sequence of Function Calls with Intermediate Results
```{webr-r}
# Compute the population standard deviation
x <- c(1, 2, 3, 4, 5)
# Step 1: Compute the mean of x
mean_x <- mean(x)
# Step 2: Compute the squared deviations from the mean
squared_deviations <- (x - mean_x)^2
# Step 3: Compute the mean of the squared deviations
mean_squared_deviations <- mean(squared_deviations)
# Step 4: Take the square root to get the population standard deviation
population_sd <- sqrt(mean_squared_deviations)
print(population_sd)
#> [1] 1.414214
```Piping
Offers improved readability and maintainability by breaking down complex operations into a sequence of simple steps.
introduced in the magrittr package and later incorporated into dplyr.
The pipe operator %>% passes the result of one function call as the first argument to the next function.
In this example:
data %>% starts the pipeline with the data.
-(mean(.)) subtracts the mean of data from each element.
^(2) squares each deviation.
mean() computes the mean of the squared deviations.
sqrt() takes the square root of the variance to get the standard deviation.
Lexical Scoping
Scoping determines how values are found in an R environment when a name is referenced.
Lexical scoping, in particular, refers to how R resolves the value of a variable name based on where the variable is defined in the source code, rather than where it is called.
 Explanation
Explanation
In this example, we have a variable x defined in the global environment and another variable x defined inside the function f(). When we call f(), the function returns the value of x defined within its own scope, which is 20.
Global Environment:
x <- 10: A variable x is assigned the value 10.Function Definition:
f <- function() { ... }: A function f is defined. Inside this function:
x <- 20: A new variable x is assigned the value 20 within the local scope of the function.
return(x): The function returns the value of x from its local scope.
Function Call:
result <- f(): The function f() is called, and the returned value (20) is assigned to result.Output:
print(result): This prints 20 to the console.The four primary rules of R’s lexical scoping:
- **Name Masking**:
- If a variable name is defined in multiple nested environments (e.g., global environment and function's local environment), the closest (most nested) definition takes precedence. This is known as name masking.
```{webr-r}
x <- 10 # Global environment
f <- function() {
x <- 20 # Local environment of f()
print(x) # Prints 20, not the global 10
}
```
- **Functions versus Variables**:
- Functions and variables are treated similarly in scoping. They both follow lexical scoping rules, meaning their visibility is determined by their definition location in the source code.
```{webr-r}
x <- 10
g <- function() {
print(x) # Accesses the global variable x
}
```
- **A Fresh Start**:
- Each function call creates a new local environment with a fresh set of bindings (variables). This means each function call operates with its own set of variables that are independent of other function calls.
```{webr-r}
x <- 10
h <- function() {
x <- 20
i <- function() {
print(x) # Accesses x from the local environment of h()
}
i()
}
```
- **Dynamic Lookup**:
- Variables are looked up dynamically based on the scope hierarchy at the time of execution, not at the time of definition. This means that a variable's value is determined by the environment it is currently being accessed from, not where it was defined.
```{webr-r}
x <- 10
j <- function() {
print(x) # Accesses the global variable x at runtime
}
```Lazy evaluation
Lazy evaluation refers to the behavior where function arguments are not evaluated until they are actually needed or accessed within the function’s body.
This concept ensures efficiency by delaying computation until necessary.
 In this example:
In this example:
The function f is defined to accept an argument x.
Inside f, x is not used in any computation or operation.
When f is called with f(x = 10), the argument x is passed but never accessed within the function’s body.
Despite x being provided as an argument when calling f, no error occurs because R does not evaluate x unless it is explicitly used within the function. This demonstrates lazy evaluation: R postpones the evaluation of x until it is actually needed inside the function.
Promises
Promises are integral to lazy evaluation.
They encapsulate the expression and environment where an expression should be evaluated, deferring its computation until its value is explicitly needed.
 Explanation:
Explanation:
Promise Creation: In function f, x + y forms a promise. It represents the expression to be evaluated (x + y) and the environment where it should be evaluated (the environment where f is called).
Deferred Evaluation: When g calls f(z, 5), z and 5 are passed as arguments to f. However, x + y (the promise) is not immediately evaluated. Instead, it remains as a promise until its value is explicitly needed.
Eager vs Lazy Evaluation: R will evaluate the promise (compute x + y) only when the result of f(z, 5) is actually required, such as when result is assigned the value returned by g(6).
Environment Sensitivity: Promises are evaluated in the environment where the function is called (g in this case), ensuring that the correct values of variables (z and 5) are used at evaluation time.Missing
Missing Argument
The missing() function is useful for determining whether an argument passed to a function is explicitly provided by the user or if it defaults to a predefined value within the function definition.
 Explanation:
Explanation:
Function Definition: my_function is defined with x having a default value of 10.
Using missing(): Inside my_function, missing(x) checks if x was provided explicitly by the user or if it defaults to 10.
Calling my_function():
When called without arguments (my_function()), x defaults to 10. missing(x) returns TRUE because x is not provided by the user.
Therefore, the function prints a message indicating that the default value of x is being used.
Calling my_function(20):
Here, x is explicitly provided as 20. missing(x) returns FALSE because x is provided by the user.
The function prints a message showing that x was indeed provided by the user with the value 20.
The sample() function in R requires at least two arguments:
x: This argument specifies the vector or set of values from which to sample.
size: This argument specifies the number of samples to draw from x.
There is also an optional argument:
replace = FALSE).The %||% infix function in R is typically used to simplify expressions where you want to use a default value or fallback to an alternative if the left-hand side is NULL.
This can be particularly useful in defining default arguments or handling optional parameters in functions.
 Explanation:
Explanation:
Definition of %||%: Defines a custom infix function %||% that checks if x is NULL or not. If x is not NULL, it returns x; otherwise, it returns default.
Redefined sample2 function: Uses %||% to provide default values for size and replace parameters:
size %||% 1: If size is not provided (NULL), default to 1.
replace %||% FALSE: If replace is not provided (NULL), default to FALSE.
Examples:
sample2(1:10): Uses default size = 1 and replace = FALSE.
sample2(letters, size = 3, replace = TRUE): Specifies size = 3 and replace = TRUE.
sample2(LETTERS, size = 5): Uses size = 5 with default replace = FALSE.
when using %||%, the right-hand side (the default value or fallback) will only be evaluated if the left-hand side is NULL.
Exiting Function
Functions can return values either implicitly or explicitly:
Implicit Returns: Implicit return occurs when the last evaluated expression within the function is automatically returned as its result
In this example, a + b is the last expression in the function add_numbers, so its result (8) is implicitly returned when the function is called.
Explicit Returns:
Explicit return involves using the return() function to explicitly specify the value to be returned from the function. This is useful when you want to return early from a function or when the return value isn’t the last evaluated expression:
 In this example,
In this example, return(sum_numbers / length(numbers)) explicitly returns the calculated average. The return() function can be used anywhere within the function body to specify the return value.
Invisible Values:
- functions typically return values that are visible when called interactively. This means that the result of calling the function will be printed in the console or displayed in some manner.
```{webr-r}
square <- function(x) {
x^2 # Returns the square of x
}
# Calling the function and displaying the result
square(5)
# Output: 25
```
{width="76"} In this example, when `square(5)` is called, `25` is printed as output because the function `square` returns the square of its input `x`.
- sometimes you may want a function to return a value invisibly, meaning it is computed and returned as the function result but not printed or displayed by default.
- This is useful when you want to perform a computation within a function but don't want the result to clutter the console output or interfere with subsequent operations.
```{webr-r}
compute_sum <- function(a, b) {
result <- a + b
invisible(result) # Return result invisibly
}
# Calling the function and capturing the result
sum_result <- compute_sum(10, 20)
print(sum_result) # Output: 30
```
{width="78"} In this example, `compute_sum(10, 20)` calculates the sum `30`, but because `invisible(result)` is used, the result `30` is returned invisibly. This means it's computed and can be assigned to a variable (`sum_result`), but it doesn't print directly to the console unless explicitly printed or used.
- `withVisible()` is a function that allows you to return a value along with a visibility flag.
```{webr-r}
compute_product <- function(x, y) {
result <- x * y
visible_result <- withVisible(result) # Capture result with visibility flag
return(visible_result)
}
# Example usage:
prod_result <- compute_product(7, 8)
# Checking the returned object
prod_result
```The stop() function is used to terminate the execution of a function immediately and throw an error message.
Exit handlers:
Exit handlers are useful for ensuring that certain actions are performed whenever a function exits, regardless of whether it exits normally or due to an error:
 In this example:
In this example:
The modify_global_state() function modifies the global state by creating a temporary variable temp_var.
on.exit() is used to set up an exit handler that will execute regardless of how the function exits.
Inside the exit handler, you can perform cleanup actions such as removing temporary variables (rm(list = ls(pattern = "^temp_"))).
The function includes a commented-out line (stop("Error: Simulation of an error condition.")) to simulate an error condition. If this line is uncommented, the exit handler will still execute.
Prefix
```{webr-r}
# Example function with named arguments
prefix_example <- function(arg1, argument_long_name, arg3) {
cat("Argument 1:", arg1, "\n")
cat("Argument with long name:", argument_long_name, "\n")
cat("Argument 3:", arg3, "\n")
}
# Calling the function using exact names
prefix_example(arg1 = 1, argument_long_name = 2, arg3 = 3)
# Using unique prefixes (partial matching)
prefix_example(a = 1, argument_long = 2, arg3 = 3)
# Arguments by position
prefix_example(1, 2, 3)
``` In this example:
In this example:
The prefix_example function takes three arguments: arg1, argument_long_name, and arg3.
You can call prefix_example using exact names (arg1 =, argument_long_name =, arg3 =), which is straightforward and explicit.
R also supports partial matching. For instance, a = 1 matches arg1, and argument_long = 2 matches argument_long_name because it is the only argument starting with argument_long.
Lastly, you can call the function by providing arguments in the order of their definition (1 for arg1, 2 for argument_long_name, 3 for arg3).
You can create a custom infix function using % symbols:
 In this example:
In this example:
The function %^% is defined to compute the exponentiation of base raised to the power of exponent.
To define an infix function, the function name is enclosed in backticks (``), and %^% indicates the custom infix operator for exponentiation.
When you use %^%, it calculates 2^3, resulting in 8.
when defining infix functions, their names can include any sequence of characters except for %. Here’s an example illustrating how to define and use an infix function with special characters:
```{webr-r}
# Define an infix function with special characters
`%+%` <- function(x, y) {
paste(x, "+", y)
}
# Using the infix function
result <- "Hello" %+% "World"
print(result) # Outputs: "Hello + World"
``` In this example:
In this example:
The function %+% is defined to concatenate two strings x and y with a “+” sign in between.
The function name %+% is enclosed in backticks (``) to indicate it’s an infix operator.
When calling the infix function, such as "Hello" %+% "World", you don’t need to escape special characters like + within the function call itself.
infix operators follow default precedence rules where they are composed from left to right. This means that when multiple infix operators are used in an expression, they are evaluated based on their position and precedence.
Replacement functions are denoted by their special name format xxx<-, where xxx is the name of the function they replace.
These functions modify their arguments in place by assigning a new value.
They must have arguments named x (the object to be modified) and value (the new value to assign), and they should return the modified object.
| Special_Form | Prefix_Form |
|---|---|
| `(`(x) | (`(x) |
| `{`(x) | {`(x) |
| `[`(x, i) | [`(x, i) |
| `[[`(x, i) | [[`(x, i) |
| `if`(cond, true) | if (cond) true |
| `if`(cond, true, false) | if (cond) true else false |
| `for`(var, seq, action) | for(var in seq) action |
| `while`(cond, action) | while(cond) action |
| `repeat`(expr) | repeat(expr) |
| `next`() | next() |
| `break`() | break() |
| `function`(alist(arg1, arg2), body, env) | function(arg1, arg2) {body} |
Data Wrangling in a data science project typically follows the stages illustrated(Wickham and Bryan 2023) below:
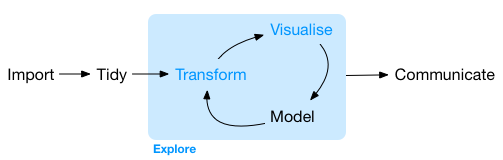
Import Data
Importing data is crucial and typically involves fetching data from files, databases, or web APIs and loading it into an object within R.
Tidy Data
Imported data into R, must be tidied into a consistent format where each column represents a variable and each row corresponds to a unique observation.
Transformation
Transformation focuses on specific observations, creating new variables based on existing ones, and deriving summary statistics.
Visualization
Visualization often uncovers unexpected or hidden patterns and prompts new questions about the data. This can help refine your questions or indicate the need for additional data.
Modeling
Once you have precisely refined your questions, you can employ models to answer them. Modeling helps in making predictions and understanding the relationships within your data.
Communication
The final and crucial step is to effectively communicate your findings to others. Clear communication ensures that your insights are understood and can be acted upon by your audience.